First observation of unusual DNA structures in archaea
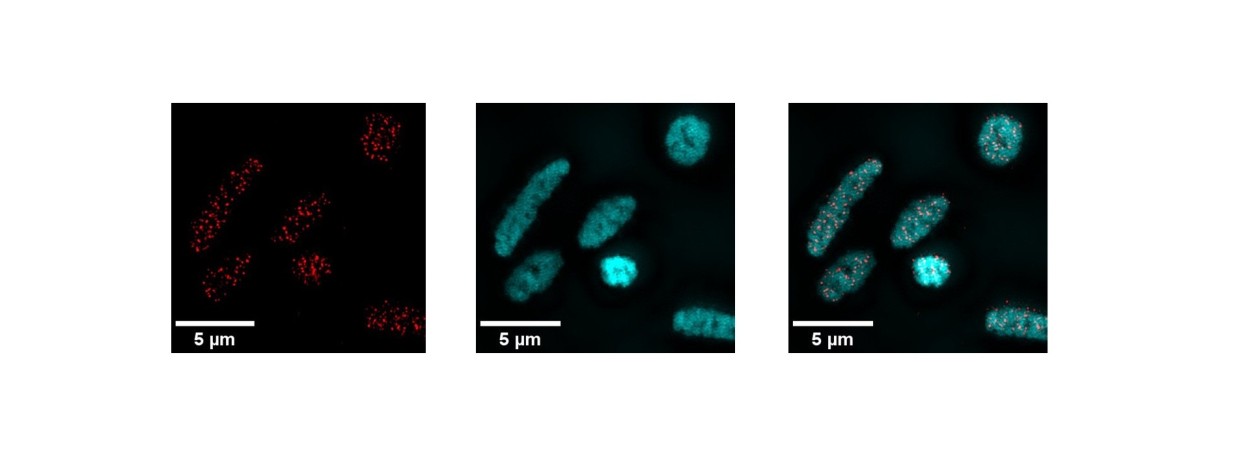 Detection in super-resolution microscopy, in Haloferax volcanii, of G-quadruplex structures (red), DNA (cyan), and their superposition.
Detection in super-resolution microscopy, in Haloferax volcanii, of G-quadruplex structures (red), DNA (cyan), and their superposition.
The double helix is famous as a symbol of the genome. It is indeed in this form that DNA is found in the cells of living beings. At least, for the most part. Behind this double helix norm lies a diversity of unusual structures of nucleic acid strands (the famous bases A, C, G, T). Scientists estimate that around 13% of the genome of a human cell could be assembled differently.
G-quadruplexes in particular are one such alternative. They are formed of guanine (G) bases, which assemble in groups of four according to a specific plan. “These planes can stack up and create different DNA molecule configurations, with four strands, two strands, or even a single strand,” explains Lionel Guittat, a professor and researcher at Université Sorbonne Paris Nord and member of the LOB.
Super-resolution microscopy
“A lot of work has already been done to identify G-quadruplexes in eukaryotic cells such as those found in mammals, but nothing has been done on archaea,” says the biologist. Yet archaea are very interesting. They are single-celled organisms, like bacteria, but recent genomic studies (particularly thanks to massive sequencing) have revealed that eukaryotes are more closely related to archaea than to bacteria. It seems that eukaryotes originated from a fusion between an archaea and a bacteria.
Lionel Guittat and his team, including Kate Sorg and Zackie Aktary, have identified G-quadruplexes (G4) in archaea for the first time. More specifically, in the species Haloferax volcanii, cultivated at the LOB.
First, an identification algorithm, based on the sequenced genome of the organism, analyzed the base sequence for potential G4 sites and identified more than 5,000. Most importantly, the scientists were able to visualize them. To do this, antibodies specifically known to be able to attach to G4s were introduced into Haloferax volcanii cells. This antibody was itself linked to a fluorescent molecule, meaning that it glows when illuminated in the right way.
Using super-resolution fluorescence microscopy (SIM, Structured Illuminated Microscopy), capable of distinguishing details down to 100 nanometers, the team was able to see the G4s within the archaea. “We carried out several checks to understand exactly what we were observing. In particular, if we add a protein that degrades DNA, we no longer see a signal. This means that our experiment clearly highlights DNA structures,” emphasizes Lionel Guittat. Another experiment shows that G4s are also present on the RNA of archaea.
Understanding the function of G4s
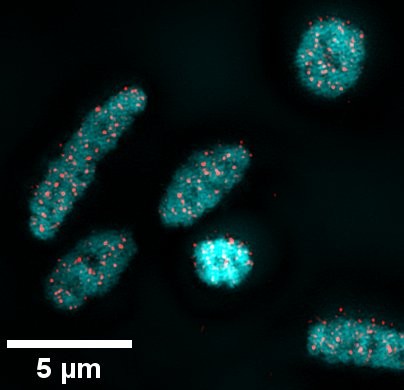
By taking the study further using STORM (Stochastic Optical Reconstruction Microscopy) super-resolution imaging, Kate Sorg was able to quantify more precisely the number of G4s that can be detected in a Haloferax Volcanii cell: around fifty.
These results were also confirmed in another archaea species, Thermococcus barophilus, in collaboration with the RNA Biology in Archaea team (MCD – CBI) at the University of Toulouse. Following this initial breakthrough, the researchers are already working on the next step: identifying the precise sequences in the archaeal genome where the observed G4s are located and learning more about their function. “We know that G4s can inhibit or, conversely, activate DNA replication, its transcription into RNA, or even translation into proteins, but this remains to be understood, particularly in archaea,” says Lionel Guittat.
Reference :
Zackie Aktary, Kate Sorg et al., Archaeal G-quadruplexes: a novel model for understanding unusual DNA/RNA structures across the tree of life, Nucleic Acids Research, Volume 54, Issue 4, 27 February 2026, gkag067, https://doi.org/10.1093/nar/gkag067
*LOB: a joint research unit CNRS, Inserm, École Polytechnique, Institut Polytechnique de Paris, 91120 Palaiseau, France
 Support l'X
Support l'X